Article
Anomalous subdiffusive measurements by fluorescence correlations spectroscopy and simulations of translational diffusive behavior in live cells
1Mathematics Department, German University in Cairo, New Cairo City; 2Laboratory of Molecular Cell Dynamics, Faculty of Advanced Life Sciences, Hokkaido University, Sapporo; 3Helios Clinical Center of Emergency Medicine, Department of Internal Medicine, 51688 Koeln-Wipperfuerth, Germany
Competing Interests: The authors have declared that no competing interests exist.
Received December 26, 2013; Revision received March 01, 2014; Accepted March 2, 2014; Published March 15, 2014
Abstract Using a combination of optical experiments and computer simulations, we found that live cells act as traps to produce anomalous subdiffusive translational motion of molecules exemplified by glucocorticoid receptor α. Glucocorticoid receptor α was expressed as a fusion protein with the green fluorescent protein in living mammalian U2OS cells. The measurements were carried out by fluorescence correlation spectroscopy. The GFP-tagged glucocorticoid receptor α was either a homodimer or a monomer in the nucleoplasm depending on the presence or absence of the stimulus dexamethasone. Our simulations showed that the experimentally measured rates of translational diffusion could be attributed to spatial and temporal traps. Thus, traps acted by spatial and temporal heterogeneity and randomness, respectively. As we prove here for the first time, that anomalous translational diffusion caused by spatial randomness was different from anomalous translational diffusion caused by heterogeneous temporal randomness. For this purpose, we explored two classes of transformations in the time domain for which the assumption that they are stable and have probability density functions was satisfied. The first one was the Inverse Gamma distribution and the second class was a stable Levy distribution. The spatial exponent α that was extracted from time series expressed scale invariance in the space domain. The time exponent γ measured limited time scaling of the embedding complex dynamics. Hence, both exponents must not be mixed up by a single exponent. Fluorescence correlation spectroscopy is the method of choice for measuring the exponents of anomalous subdiffusive translational motion of molecules in live cells.
Keywords: Anomalous translational diffusion, fluorescence correlation spectroscopy, live cells, simulation, subdiffusive transport
Introduction
Anomalous translational diffusion and subdiffusion, respectively, is a breakdown of the laws of mass action [1]. As opposed to normal translational diffusion, in which the movement of molecules is not correlated with their previous position, anomalous translational diffusion molecules are spatially and temporally correlated [1-3]. This spatio-temporal correlation reflects a fundamentally different behavior compared with the one in dilute or very dilute solution which, for example, affects the spread of molecules within live cells.
Translational diffusion coefficients typically are calculated by comparing the relative spread of a fluorescent molecule over time points [4], or by measuring the time constant of recovery after photobleaching [5]. Fluorescence correlation spectroscopy (FCS) offers an extremely powerful and successful technique for dealing with molecular interaction in dilute and very dilute solutions by delivering constant translational diffusion coefficients [4, 6-27]). Until now, characterization of translational diffusion in live cells has relied almost exclusively on measurements of constant translational diffusion coefficients [28, 29]. There is some evidence that in live cells the apparent translational diffusion coefficients may not be constant, but instead can vary over time, even for inert molecules [30-34]. The dynamics was observed by FCS. However, those measurements and theory only considered a single “dynamic” exponent [30-34]. For the first time, we have developed the theoretical formulation and an FCS-based method in order to separate the spatial subdiffusion from temporal subdiffusion [1, 2]. We have observed that FCS is superior at calculating spatially and temporally variable diffusion coefficients and our stimulated and experimental data support this.
Trapping of molecules in live cells can exert at least two types of effects that could be important for function. First, live cells will have a strong influence on how far and how fast locally produced molecules will travel within the cytoplasm or the nucleoplasm. Over a time scale of 1 s, a molecule with an apparent diffusion coefficient Dapp of 2 μm2/ms, would diffuse about 60 μm. Thus, as long as molecules remained within the cytoplasm, their movement could be dominated by an anomalous diffusion process over distances. Second, anomalous diffusion reflects an increase in the spatial and temporal correlation of diffusing molecules which would be expected to promote activation of biochemical networks by intracellular signals trapped in live cells [1, 35, 36].
Traps act by spatial and temporal heterogeneity and randomness, respectively. The amount of heterogeneity (irregularity, uncertainty) in trajectories of dynamical systems can be quantified in various ways. From a geometrical point of view, exponents measure the dependence of the future behavior on small changes in the system’s initial conditions. A mathematician might use entropy as a measure of the system’s heterogeneity in predicting the future from its past. Extensions of random walk models beyond normal Brownian motion, for example, to processes governed by steps or waiting times with heavy-tailed distributions are used to describe heterogeneities in live cells [1, 2]. At the single-molecule level, chemical and physical processes are stochastic, meaning that a reaction takes a variable time to complete. The distribution of waiting times contains information about the mechanism of the process, resulting in anomalous translational diffusion behavior of molecules that are not absorbed on or firmly bound to (i.e., immobilized on) intracellular structures. Anomalous translational diffusion allows new questions to be addressed on how to decide from a subset of single molecules how heterogeneous the system is in time. The important distinction we make is between space (structure)-dependent and time (rate)-dependent heterogeneity of anomalous translational diffusion of freely diffusing macromolecules [1, 2], with the exponent α for spatial heterogeneity and the exponent γ for temporal heterogeneity [1, 2]. To our best knowledge, experiments have never measured a value α solely due to cellular crowding.
For the first time, Klafter et al. [37] and Foldes-Papp et al. [1] proposed independently and simultaneously that subdiffusive mechanisms of mixed origins can coexist. Complex situations, e.g. when a particle moves in a percolation structure, in a tortuous channel, if it is a part of the large molecule, or if it gets trapped [38], lead to subdiffusion of mixed origins [37], which is indeed observed experimentally [2, 1, 39]. This new physical concept of anomalous translational diffusion is problematic, but it is verified in this paper. We separate both exponents α and γ in experiments and simulations and examine the heterogeneity of trajectories produced by interval transformations. Let an interval transformation 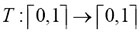 be called the continuous function for the time steps in the continuous waiting time distribution (CTRW) of the 3D random walk that is performed on a generalized Sierpinski carpet (underlying fractal structure). The transformation T denotes that it is just the inverse Gamma distribution or a stable Levy distribution of the variable time (Table 2). In the simulations, there are a finite number of time intervals (time steps) such that the interval transformation is monotone and continuous. Thus, the interval transformation preserves the given order of time steps. The focus of this paper is to show that the martingale
be called the continuous function for the time steps in the continuous waiting time distribution (CTRW) of the 3D random walk that is performed on a generalized Sierpinski carpet (underlying fractal structure). The transformation T denotes that it is just the inverse Gamma distribution or a stable Levy distribution of the variable time (Table 2). In the simulations, there are a finite number of time intervals (time steps) such that the interval transformation is monotone and continuous. Thus, the interval transformation preserves the given order of time steps. The focus of this paper is to show that the martingale 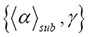 is bounded (see Fig. 3). The martingale is defined by a sequence of random variables of the exponents
is bounded (see Fig. 3). The martingale is defined by a sequence of random variables of the exponents 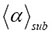 and γ for which the expectation of the next value in the sequence is equal to the present observed value. In order to avoid frequent normalizing by logarithms, we henceforth take all logarithms to base e,
and γ for which the expectation of the next value in the sequence is equal to the present observed value. In order to avoid frequent normalizing by logarithms, we henceforth take all logarithms to base e,
Using a combination of optical experiments and computer simulations, we consider the glucocorticoid receptor α (GRα). GRα is a ligand-regulated transcription factor, a member of the nuclear receptor superfamily that regulates a variety of physiological functions. It is widely thought that unliganded GRα primarily localizes in the cytoplasm as part of a multiprotein complex that includes chaperone proteins and immunophilins [40-42]. Upon ligand binding, GRα is translocated to the nucleoplasm in the nucleus, where it works either as a homodimer that binds to positive or negative glucocorticoid response elements (GRE) located in the promoter regions of target genes or as a monomer that cooperates with other transcription factors to induce transcription [43-46]. In this article, we show that GRα is either a homodimer or a monomer in the nucleus depending on the presence or absence of the stimulus dexamethasone.
The same acronym is used for fluorescence fluctuation spectroscopy and fluorescence correlation spectroscopy (FCS). These terms have the same meanings and, therefore, we use the same
terminology throughout the manuscript, for example, fluorescence correlation spectroscopy.
terminology throughout the manuscript, for example, fluorescence correlation spectroscopy.
Basic theory of anomalous subdiffusive motion (transport) in live cells
In uniform Euclidean systems, the mean-square displacement MSD of a random walker is proportional to the time t for any number of spatial dimensions d. However, in disordered systems, this physical law is not valid in general, and the diffusive law becomes anomalous and subdiffusive, respectively, as given by Eqn. 1 [1, 2, 37-39].
A linear dependence, i.e. α =1, is a fingerprint of normal translational diffusion in dilute or very dilute solution. Any other value of α corresponds to anomalous translational diffusion. Here, we focus on subdiffusion, i.e. 0 < α < 1. Subdiffusive motion can appear due to geometric and/or energetic disorders [2]. An example of geometric restraints is molecular crowding in living cells.
In order to take account of temporal randomness of molecular interaction, i.e. time (rate)-dependent sources of anomalous translational diffusion behavior, we performed the 3D random walks on fractal support as a continuous time random walk (CTRW). The diffusive law becomes
The new and important feature of Eqn, 2 is that the spatial and temporal coordinates are decoupled [1]. The exponent α now quantifies the crowding conditions and the exponent γ the temporal heterogeneity.
Fluorescence-based techniques, such as fluorescence correlation spectroscopy (FCS) enable protein-protein interactions to be spatially and temporally resolved in live cells [22, 29]. The auto- and dual color cross-correlation functions for fluorescence intensity fluctuations [24, 26] can be written in terms of subdiffsion as follows [2, 1]:
where 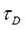 is the translational diffusion time, s the geometric parameter of the laser focus, dim = 3 stands for 3-dimensional measurements in the cytoplasm or the nucleus of live cells, and τ is the correlation time of the fluorescent molecules (time scale of diffusive motion). DC is the limiting value of G(τ) for
is the translational diffusion time, s the geometric parameter of the laser focus, dim = 3 stands for 3-dimensional measurements in the cytoplasm or the nucleus of live cells, and τ is the correlation time of the fluorescent molecules (time scale of diffusive motion). DC is the limiting value of G(τ) for 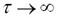 , which is usually 1 for the normalized fluorescence intensity function G(τ). We get α when we fit Eqn. (3) to time traces that are recorded for a subpopulation of single molecules of the same kind without interacting partner, e.g. without ligand, in the crowded environment of a living cell and its cellular compartments. Knowing the crowding parameter α for the cell type as well as the cellular compartment, the heterogeneous parameter γ can be extracted from the measurements in the presence of the interacting reaction partner, e.g. ligand, for the same (fixed) α value.
, which is usually 1 for the normalized fluorescence intensity function G(τ). We get α when we fit Eqn. (3) to time traces that are recorded for a subpopulation of single molecules of the same kind without interacting partner, e.g. without ligand, in the crowded environment of a living cell and its cellular compartments. Knowing the crowding parameter α for the cell type as well as the cellular compartment, the heterogeneous parameter γ can be extracted from the measurements in the presence of the interacting reaction partner, e.g. ligand, for the same (fixed) α value.
Materials and Methods
Chemicals and plasmids
Dexamethasone (Sigma) was prepared as 100 µM ethanol solutions, and diluted with Opti-MEM (GIBCO) to 10 µM just before use. Three µL of the ethanol/Opti-MEM solution were added to 300 µL of culture medium. The construction of the plasmids used is described elsewhere [41].
Cell culture, transient transfection and preparation of cell extracts
Mammalian U2OS cells do not express endogenous GRα. U2OS cells were obtained from ATCC, and maintained in a 5% CO2 humidified atmosphere at 37°C in McCoy’s 5A Medium (GIBCO) supplemented with 10% Charcoal Stripped FBS (GIBCO). Cells were subcultured every 3 or 4 days. A day before transient transfection, cells were plated on LAB-TEK 8-wells chambered coverslips (Nalge Nunc International) to provide 10000 cells per wells, and were transfected using Optifect (Invitrogen) with 100 ng/well pEGFP (monomeric GFP) or pEGFP-hGRα (monomeric GFP fusion with the human GRα) according to the manufacturer’s instructions.
FCS measurements
Fluorescence correlations spectroscopy (FCS) also called fluorescence fluctuation spectroscopy [12, 13, 15, 16, 24] is traditionally a solution-phase, optical methodology that uses a very thin laser beam to detect the random Brownian motion of a fluorescent-labeled molecule in a tiny volume of solution (or membrane) no bigger than the laser beam aimed through it (about one femtoliter) [22, 24, 26]. The FCS analysis is performed by using correlation functions like the autocorrelation function of fluorescence intensity fluctuations or the dual color cross-correlation function of fluorescence intensity functions in two colors that is mathematically of the same type as the autocorrelation function ([24, 26], and references cited therein), although other approaches have been proposed and are used. The field was started by Magde, Elson and Webb in 1972 [47]. Eigen and Rigler first resolved technical and conceptual limitations of FCS by using confocal pinholes [48]. Today, FCS for recording fluorescence fluctuations at single points is a mature and sophisticated technology.
The FCS measurements in the cell nucleus were performed with a LSM710-ConfoCor3 system (manufactured by Carl Zeiss Inc.) before and 30 minutes after addition of dexamethasone. Each set of FCS measurements was carried out five times with a measurement time of 15 s.
FCS data
Transfection of U2OS cells with monomeric GFP and subsequent FCS measurements in the nucleus yielded the crowding exponent without any interaction part: α = 0.87958 ± 0.0044 (all diffusive time scales), α = 0.95294 ± 0.01027 (diffusive time scale 0.1 - 1 ms), α = 0.90001 ± 0.03913 (diffusive time scale 1 - 10 ms). The 0.01 - 0.1 ms timescale is too small to extract reliable values for the anomalous motion.
The constructed plasmid of the monomeric GFP fusion with the GRα (pEGFP-hGRα, GRα-GFP) was transiently transfected into U2OS cell and is expressed in the nucleus. The subsequent FCS measurements were performed in the nucleus of the U2OS cell without dexamethasone stimulus (DEXminus) and after dexamethasone stimulus (DEXplus).
Simulations
The approaches that we use are random walks on a three-dimensional (3D) fractal support and continuous time random walks (CTRW) [37, 49]. Both approaches aim on the clarification of the interrelation of random processes connected with real translational diffusion phenomena observed in experiments. The majority of work found in literature, however, is restricted to one dimensional or two dimensional systems [37, 50]. Only a few articles deal with the real and three-dimensional problem [51].
Our algorithms described in detail in refs. [1, 2] treat every walker as a single unit that performs a random walk over the 3D Sierpinski carpet (generalized three-dimensional Sierpinski carpet). It is a random walk on a fractal 3D structure (RWF). The movement of each individual walker at each time step is controlled, independently for each degree of freedom, by a step size and by a probability distribution Pn(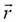 ) with probabilities that determine in which direction the walker takes a step, i.e. how the steps are distributed. In a simple discrete random walk, the walker advances one step in unit time. Each step is taken to a nearest neighbor of the site. The simple random walker can step with equal probability to any of the nearest neighbor sites that belong to the grid. There are several ways of assigning transition probabilities for stepping from site to site. The motion of the Brownian molecule is generated with a random number generator delivering pseudo-random numbers used for the steps in all three spatial directions. We generate a random Brownian walk by randomly selecting steps in the three coordinate directions. The three coordinate directions are generated by a permutation (for details, see ref. [1]).
) with probabilities that determine in which direction the walker takes a step, i.e. how the steps are distributed. In a simple discrete random walk, the walker advances one step in unit time. Each step is taken to a nearest neighbor of the site. The simple random walker can step with equal probability to any of the nearest neighbor sites that belong to the grid. There are several ways of assigning transition probabilities for stepping from site to site. The motion of the Brownian molecule is generated with a random number generator delivering pseudo-random numbers used for the steps in all three spatial directions. We generate a random Brownian walk by randomly selecting steps in the three coordinate directions. The three coordinate directions are generated by a permutation (for details, see ref. [1]).
In addition, in a three-dimensional (3D) RWF with a continuous waiting time distribution (CTRW), the molecule has to wait for a time on each site of the fractal before performing the next step. This waiting time is a random variable independently chosen at each new step according to a continuous distribution. The waiting time distribution is independent of the location of the molecule, i.e. we decouple spatial and temporal coordinates. One can think about this process as a diffusive motion among traps, but the trapping time will change for each site on the lattice if this site is visited again during the random walk. In our case, the Inverse Gamma distribution and a stable Levy distribution are used as waiting time distributions for which the assumption that they are stable and have probability density functions was satisfied. The probability to perform n steps during time t is denoted by χn(t), which is related to the waiting time-distribution by the Laplace transform (for details, see ref. [1]). This probability χn(t) is needed to analyze the MSD for a random walk on a fractal carried out as a CTRW.
The finger print of RWF is its mean square displacement (MSD). The MSDs on 3D random walks on the fractal support of a generalized Sierpinski carpet are shown in Figures 1 and 2. This distribution of scaling exponents renders a system inhomogeneous: an ensemble of simple diffusers with different translational diffusion coefficients at a fixed time exponent γ =0.81398 (Fig. 1) and a fixed time exponent γ=0.60497 (Fig. 2). Since α and γ satisfy 0 < α, γ < 1, so does the product α times γ and the diffusive motion is thus subdiffusive.
Results and Discussion
Mammalian U2OS cells do not express endogenous GRα. The FCS measurements of the monomeric GFP expressed in the nucleus of U2OS cells yielded the crowding exponent α in the nucleus without any interaction part: α = 0.87958 ± 0.0044 (all diffusive time scales). We fitted Eqn. 3 for the parameter α for a population of single molecules without interaction partner/ligand. The measured FCS parameters of GRα-GFP in the nucleus of mammalian U2OS cells are depicted in Table 1. We controlled the experiments to ensure the existence or the lack of the reaction partner dexamethasone. The parameter γ was extracted from the measurements in the absence and presence of the interacting reaction partner dexamethasone. Over all diffusive time scales, we obtained in this way a good guess for the crowding exponent α and the time exponent γ.
Figure 1. Simulations of MSD(t) versus t of 500 tracks (sample size) in an observation volume of ΔV= 0.2 fL. The simulations were performed with different geometric fractional structure α at a fixed time exponent γ =0.81398 that is the measured value of GRα-GFP in the nucleus of mammalian U2OS cells found before stimulus with dexamethasone. The resulting mean scaling exponent 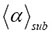 is listed on top of the plots. The α value, which was used in the generation of the stochastic tracks and from which the value
is listed on top of the plots. The α value, which was used in the generation of the stochastic tracks and from which the value 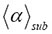 was derived, is also given on top of each plot. All calculations were performed by three-dimensional RWF with the inverse gamma function as CTRW (see Table 2). For detailed explanation see main text.
was derived, is also given on top of each plot. All calculations were performed by three-dimensional RWF with the inverse gamma function as CTRW (see Table 2). For detailed explanation see main text.
Figure 2. Simulations of MSD(t) versus t of 500 tracks (sample size) in an observation volume of ΔV= 0.2 fL. The simulations were performed with different geometric fractional structure α at a fixed time exponent γ=0.60497 that is the measured value of the GRα-GFP in the nucleus of mammalian U2OS cells found after stimulus with dexamethasone. The resulting mean scaling exponent 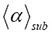 is listed on top of the plots. The α value, which was used in the generation of the stochastic tracks and from which the value
is listed on top of the plots. The α value, which was used in the generation of the stochastic tracks and from which the value 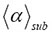 was derived, is also given on top of each plot. All calculations were performed by three-dimensional RWF with the inverse gamma function as CTRW (see Table 2). For detailed explanation see main text.
was derived, is also given on top of each plot. All calculations were performed by three-dimensional RWF with the inverse gamma function as CTRW (see Table 2). For detailed explanation see main text.
The experimentally measured exponents α and γ of Table 1 are averaged over many single molecules of the same kind. We have shown before [1] that the ergodicity is either unbroken or it is indistinguishable from broken ergodicity under such condition at the single-molecule level. The origin of the slow and intracellular translational diffusion of GR in the nucleus is still in discussion, but some part of diffusion is involved from interactions with chromatin [52, 53]. We next used simulations of subdiffusive, i.e. anomalous, intracellular translational diffusion to understand how space and time heterogeneity might slow down the translational diffusive transport of a macromolecule such as GRα-GFP in the nucleus of mammalian U2OS cells. As given in Table 2, our simulations were based on the conditions of the measurements summarized in Table 1.
Table 1. Measured FCS parameters in the nucleus of mammalian U2OS cells that expressed the fusion protein of GRα-GFP. The rate of translational diffusion is expressed by the translational diffusion time τD in the absence and presence of dexamethasone. f stands for fixed values in the FCS data fits. The parameter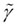 is given by
is given by 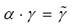 . For explanations, see main text.
. For explanations, see main text.
The plots of simulated mean-square displacement MSD(t) versus t are depicted in Figures 1 and 2. 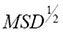 is the mean distance from the starting point, e.g. in μm, that GRα-GFP in the nucleus of mammalian U2OS cells will have diffused in time, t. t is given in time steps covering a range from about 10 μs to 100 ms. The simulations of Figures 1 and 2 were performed at a fixed time exponent γ=0.81398 that is the measured value of the GRα-GFP in the nucleus of mammalian U2OS cells found before stimulus with dexamethasone (Fig. 1) and at a fixed time exponent γ=0.60497 that is the measured value of GRα-GFP in the nucleus of mammalian U2OS cells found after stimulus with dexamethasone (Fig. 2). Since
is the mean distance from the starting point, e.g. in μm, that GRα-GFP in the nucleus of mammalian U2OS cells will have diffused in time, t. t is given in time steps covering a range from about 10 μs to 100 ms. The simulations of Figures 1 and 2 were performed at a fixed time exponent γ=0.81398 that is the measured value of the GRα-GFP in the nucleus of mammalian U2OS cells found before stimulus with dexamethasone (Fig. 1) and at a fixed time exponent γ=0.60497 that is the measured value of GRα-GFP in the nucleus of mammalian U2OS cells found after stimulus with dexamethasone (Fig. 2). Since 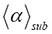 results from an averaging process that is defined as minimization (see equation (12) in ref. [2]), we need reference values α to be able to estimate the variation of
results from an averaging process that is defined as minimization (see equation (12) in ref. [2]), we need reference values α to be able to estimate the variation of 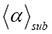 . The properly chosen α value is given on top of each plot in Figures 1 and 2. The values of α are consistent with experimental results. For a detailed discussion of the parameters and their relation among each other see ref. [2]. In order to physically interpret the experimental results of Figures 1 and 2, we shall show that the molecular system is invariant under transformation T for the time steps in the continuous waiting time distribution (CTRW) of the 3D random walk performed on a generalized Sierpinski carpet (underlying fractal structure).
. The properly chosen α value is given on top of each plot in Figures 1 and 2. The values of α are consistent with experimental results. For a detailed discussion of the parameters and their relation among each other see ref. [2]. In order to physically interpret the experimental results of Figures 1 and 2, we shall show that the molecular system is invariant under transformation T for the time steps in the continuous waiting time distribution (CTRW) of the 3D random walk performed on a generalized Sierpinski carpet (underlying fractal structure).
Let us now formulate the kind of problem we tackle here. A subdiffusive path of a molecule can make an up-move, another path makes a transverse move. One may compute independent realizations of a random walk where the following unknown quantities of paths occur: the number of up-moves, the number of transverse moves, the number of shared up-moves, the number of shared transverse moves, the number of simultaneous moves with the same starting point of which one is up and the other is transverse, and so forth. This projection reflects the fact that nearby trajectories of molecules do not necessarily arrive at the same space point at the same time. Are these realizations of a random walk equal, and if not can they be evaluated? The problem is verified as follows.
In Figure 3, we used two different continuous functions for the distribution of time steps at which the various combinations of events (moves) occur in the realizations of a random walk of molecules, namely a stable Levy distribution and the inverse Gamma distribution. There is no evidence that the two distributions (Levy distribution and inverse Gamma distribution) used in the simulations behave differently. We found that 3D single-molecule trajectories are represented by a stochastic process for martingale limits, i.e. upper and lower boundary values of the mean crowding exponent 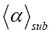 and the temporal heterogeneity exponent γ.
and the temporal heterogeneity exponent γ. 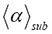 fluctuates in an intrinsically irregular and erratic manner, and grows endogenously like a slow varying function
fluctuates in an intrinsically irregular and erratic manner, and grows endogenously like a slow varying function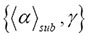 within those boundary values. In the continuous-time martingale with respect to the stochastic function
within those boundary values. In the continuous-time martingale with respect to the stochastic function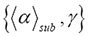 , it is a stochastic process such that the expectation of the next value in the sequence
, it is a stochastic process such that the expectation of the next value in the sequence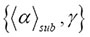 is equal to the present observed value even when all the values observed previously are known. The upper bound of the crowding
is equal to the present observed value even when all the values observed previously are known. The upper bound of the crowding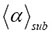 represented in the data of Figure 3 by the different generators g_12, g_18, and g_24 [1], increases if the geometric fractal becomes more dense. There is also a change in the slope of
represented in the data of Figure 3 by the different generators g_12, g_18, and g_24 [1], increases if the geometric fractal becomes more dense. There is also a change in the slope of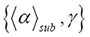 for smaller values of γ when the graph starts compared to greater values of γ when the graph ends. In fact, we have scaling in the time domain and we have scaling in the spatial domain. However, the averaging of the spatial exponent α implicitly uses the value of the temporal properties. What matters most are not the details of the microscopic interactions, but rather the nature of the paths along which order is propagated from one molecule to another distant molecule. Complex interactions can lead to simple results. This is particularly true for systems formed of many interacting subunits where unbroken ergodicity becomes indistinguishable from broken ergodicity [1].
for smaller values of γ when the graph starts compared to greater values of γ when the graph ends. In fact, we have scaling in the time domain and we have scaling in the spatial domain. However, the averaging of the spatial exponent α implicitly uses the value of the temporal properties. What matters most are not the details of the microscopic interactions, but rather the nature of the paths along which order is propagated from one molecule to another distant molecule. Complex interactions can lead to simple results. This is particularly true for systems formed of many interacting subunits where unbroken ergodicity becomes indistinguishable from broken ergodicity [1].
Hence, the anomalous translational subdiffusion describes spatial and heterogeneous (temporal) randomness in cellular systems as it was proposed by us in ref. [2]. Simplified in vitro FCS-models of the intracellular environment [28], in which crowding and binding are tuned independently, are not a useful tool to gain insight into molecular transport implicated in key cellular processes in live cells [46].
In the simulations, we took the measured exponents for spatial and heterogeneous (temporal) randomness, i.e. α and γ, and analyzed the mean-square displacement. After the receptor is bound to glucocorticoid (dexamethasone), the homodimer receptor-glucorticoid complex can take either of two pathways. The activated homodimer GR complex up-regulates the expression of anti-inflammatory proteins in the nucleus or it represses the expression of pro-inflammatory proteins in the cytosol by preventing the translocation of other transcription factors from the cytosol into the nucleus [54].
Our point of view [55-57, 1, 2] is something different from the work of Enrico Gratton and his colleagues on pair-correlation, which is used to detect diffusional barriers [58-61]. The method of spatial pair-correlation (pCF) is substantially different from the conventional FCS method. In pCF, diffusion is measured by the average time particles (molecules) take to travel between two points as the position of the two points is arbitrary. A laser beam is moved rapidly to different locations in a repeated pattern, e.g., a line, circle, or grid. The entire pattern is repeated in approximately 1 to 10 ms. If there is a barrier to diffusion that may called trap, the particle goes around the obstacle or passes over the barrier. The maximum of the spatial pair-correlation function (pCF) will be found at a longer time than in the absence of a barrier. By mapping the time of the maximum of the pCF for every pair of points in the image, the location and the size of the obstacles can be visualized. By fitting the series of correlation functions, the actual protein “diffusion law” can be obtained directly from imaging, in the form of a mean-square displacement vs. time-delay plot [59]. Future advances in the spatial pair correlation approach will involve the development of better detection methods and the development of techniques for three-dimensional correlations for the cellular environment [58].
Figure 3. Simulation of 500 tracks (sample size) in an observation volume of ΔV= 0.2 fL for each type of transformations (CTRW) in A, B and C. A-C. Different generators (fractal supports) of the 3D Sierpinski carpet were used (see ref. [1], Table 2 therein). α is a consequence of the chosen generator of the 3D Sierpinski carpet [1]. The less the exponent α, the higher the geometric crowding of the underlying structure of translational motion. Simulated data are depicted for different time exponents γ at different mean scaling exponents 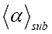 . For explanations, see main text.
. For explanations, see main text.
Many theoretical models of molecular interactions as well as biochemical and chemical reactions in dilute solution and live cells are described on the single-molecule level – although, our knowledge about the biochemical/chemical structure and dynamics mostly originates from investigations of many-molecule systems [62, 63]. At present, there are four experimental platforms to observe the movement and the behavior of single fluorescent molecules: wide-field epi-illumination, near-field optical scanning, and laser scanning confocal and multiphoton microscopy. The platforms are combined with analytical methods such as fluorescence resonance energy transfer (FRET), fluorescence auto- or dual color cross-correlation spectroscopy (FCS), fluorescence polarizing anisotropy, fluorescence quenching and fluorescence lifetime measurements. The concept of fluctuations due to the same molecule was theoretically and experimentally developed in recent years [55, 57, 8, 9, 18, 35, 36, 27, 12, 13, 62, 63, 1, 2]. Under conditions in which only one molecule is present in the observation volume being sampled, we have shown that it is possible to measure correlations due to the re-entry of the same molecule in the volume of illumination. The meaningful time [57, 55, 35, 36, 9] is the physical parameter that enables to learn some characteristic signatures of the same single molecule in solution and live cells.
Table 2. Measured exponents of anomalous translational diffusion in the nucleus of mammalian U2OS cells and their simulations.
Conclusions
A FCS-based method for the measurement of anomalous subdiffusion is described, as previously developed [1, 2]. In this original article, we used the FCS method to study the movement of GRα-GFP in U2OS cells. We also simulated diffusion with random spatial and temporal traps with the Inverse Gamma distribution and Levy distribution. The results indicate that it is possible to separate the spatial subdiffusion from temporal subdiffusion.
We found that live cells act as traps to produce anomalous subdiffusive translational motion of molecules. We have first shown here that calculations of the space exponent α from time series are possible with the use of reliable values of the time exponent γ. Averaged values of the time exponent γ have only limited abilities because γ is a measure for limited time scaling. In particular, spurious exponents γ may arise from the embedding complex dynamics and should be distinguished from the real exponents γ measured on all time scales of the complex dynamical system. Hence, both exponents must not be mixed up by a single exponent. Fluorescence correlation spectroscopy (FCS) is the method of choice for measuring the exponents of anomalous subdiffusive translational motion of molecules in live cells as demonstrated here for the first time.
Acknowledgments
Zeno Földes-Papp, who is the principal investigator, acknowledges financial support in part from the Helios Clinical Center of Emergency Medicine, Department of Internal Medicine, Koeln-Wipperfuerth, Germany, from the German University in Cairo, and from the bwGRiD Cluster Ulm that is part of the high performance computing facilities of the Federal State of Baden-Wuerttemberg (Germany), where most of the very time-consuming and expensive numerical calculations were performed. Due to space limit, we apologize for the many omissions of the pioneering work of other researchers, and mainly focus on our contributions to the field that are related to the reliable measurements of translational diffusive behavior of molecules. We do not describe here the conceptual history of FCS and related approaches.
References
- Baumann G, Place RF, Földes-Papp Z (2010) Meaningful interpretation of subdiffusive measurements in living cells (crowded environment) by fluorescence fluctuation microscopy. Curr Pharm Biotechnol 11: 527-543. [PubMed] [Google Scholar]
- Földes-Papp Z, Baumann G (2011) Fluorescence molecule counting for single-molecule studies in crowded environment of living cells without and with broken ergodicity. Curr Pharm Biotechnol 12: 824-833. [PubMed] [Google Scholar]
- Földes-Papp Z, Angerer B, Ankenbauer W, Baumann G, Birch-Hirschfeld E, et al. (1996) Modeling the dynamics of nonenzymatic and enzymatic nucleotide processes by fractal dimension. In: Losa GA, Merlini D, Nonnenmacher TF, Weibel ER, editors. Fractals in Biology and Medicine, vol 2. Boston: Birkhauser. pp. 238-254.
- Földes-Papp Z, Thyberg P, Björling S, Holmgen A, Rigler R (1997) Exonuclease degradation of DNA studied by fluorescence correlation spectroscopy. Nucleosides, Nucleotides & Nucleic Acids 16: 5-6.[Google Scholar]
- Axelrod D, Koppel DE, Schlessinger J, Elson E, Webb WW (1976) Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys J 16: 1055-1069. doi: 10.1016/S0006-3495(76)85755-4. [View Article] [PubMed] [Google Scholar]
- Rich RM, Mummert M, Gryczynski Z, Borejdo J, Sørensen TJ, et al. (2013) Elimination of autofluorescence in fluorescence correlation spectroscopy using the AzaDiOxaTriAngulenium (ADOTA) fluorophore in combination with time-correlated single-photon counting (TCSPC). Anal Bioanal Chem 405: 4887-4894. doi: 10.1007/s00216-013-6879-0. [View Article] [PubMed] [Google Scholar]
- Rich RM, Mummert M, Foldes-Papp Z, Gryczynski Z, Borejdo J, et al. (2012) Detection of hyaluronidase activity using fluorescein labeled hyaluronic acid and Fluorescence Correlation Spectroscopy. J Photochem Photobiol B 116: 7-12. doi: 10.1016/j.jphotobiol.2012.07.007. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Liao SJ, You T, Terpetschnig E, Barbieri B (2010) Confocal fluctuation spectroscopy and imaging. Curr Pharm Biotechnol 11: 639-653. [PubMed] [Google Scholar]
- Földes-Papp Z (2009) Viral chip technology in genomic medicine. In: Willard WF, Ginsburg GS, editors. Handbook of genomic and personalized medicine, vol 2. New York: Elsevier; pp. 538-561.[Google Scholar]
- Földes-Papp Z, Kinjo M, Tamura M, Birch-Hirschfeld E, Demel U, et al. (2005) A new ultrasensitive way to circumvent PCR-based allele distinction: direct probing of unamplified genomic DNA by solution-phase hybridization using two-color fluorescence cross-correlation spectroscopy. Exp Mol Pathol 78: 177-189. doi: 10.1016/j.yexmp.2005.01.005. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Costa JM, Demel U, Tilz GP, Kinjo M, et al. (2004) Specifically associated PCR products probed by coincident detection of two-color cross-correlated fluorescence intensities in human gene polymorphisms of methylene tetrahydrofolate reductase at site C677T: a novel measurement approach without follow-up mathematical analysis. Exp Mol Pathol 76: 212-218. doi: 10.1016/j.yexmp.2003.12.007. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Demel U, Tilz GP (2004) A new concept for ultrasensitive fluorescence measurements of molecules in solution and membrane: 1. Theory and a first application. J Immunol Methods 286: 1-11. doi: 10.1016/j.jim.2004.01.008. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Demel U, Tilz GP (2004) A new concept for ultrasensitive fluorescence measurements of molecules in solution and membrane: 2. The individual immune molecule. J Immunol Methods 286: 13-20. doi: 10.1016/j.jim.2004.01.007. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Kinjo M, Saito K, Kii H, Takagi T, et al. (2003) C677T single nucleotide polymorphisms of the human methylene tetrahydrofolate reductase and specific identification : a novel strategy using two-color cross-correlation fluorescence spectroscopy. Mol Diagn 7: 99-111. [PubMed] [Google Scholar]
- Földes-Papp Z, Demel U, Domej W (2002) Tilz GP (2002) A new dimension for the development of fluorescence-based assays in solution: From Physical Principles of FCS Detection to Biological Applications. Exp Biol Med 227: 291-300. [PubMed] [Google Scholar]
- Földes-Papp Z, Demel U, Tilz GP (2002) Detection of single molecules: solution-phase single-molecule fluorescence correlation spectroscopy as an ultrasensitive, rapid and reliable system for immunological investigation. J Immunol Methods 260: 117-124. [PubMed] [Google Scholar]
- Jermutus L, Kolly R, Földes-Papp Z, Hanes J, Rigler R, et al. (2002) Ligand binding of a ribosome-displayed protein detected in solution at the single molecule level by fluorescence correlation spectroscopy. Eur Biophys J 31: 179-184. doi: 10.1007/s00249-001-0204-0. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Demel U, Tilz GP (2001) Ultrasensitive detection and identification of fluorescent molecules by FCS: impact for immunobiology. Proc Natl Acad Sci U S A 98: 11509-11514. doi: 10.1073/pnas.181337998. [View Article] [PubMed] [Google Scholar]
- Stephan J, Dörre K, Brakmann S, Winkler T, Wetzel T, et al. (2001) Towards a general procedure for sequencing single DNA molecules. J Biotechnol 86: 255-267. [PubMed] [Google Scholar]
- Földes-Papp Z, Angerer B, Ankenbauer W, Rigler R (2001) Fluorescent high-density labeling of DNA: error-free substitution for a normal nucleotide. J Biotechnol 86: 237-253. [PubMed] [Google Scholar]
- Földes-Papp Z, Angerer B, Thyberg P, Hinz M, Wennmalm S, et al. (2001) Fluorescently labeled model DNA sequences for exonucleolytic sequencing. J Biotechnol 86: 203-224. [PubMed] [Google Scholar]
- Földes-Papp Z, Kinjo M (2001) Fluorescence correlation spectroscopy in nucleic acids analysis. In: Elson E, Rigler R, editors. Fluorescence correlation spectroscopy: theory and applications. Springer series in physical chemistry, vol 65. Boston: Springer. pp. 25-64. [Google Scholar]
- Lagerkvist AC, Földes-Papp Z, Persson MA, Rigler R (2001) Fluorescence correlation spectroscopy as a method for assessment of interactions between phage displaying antibodies and soluble antigen. Protein Sci 10: 1522-1528. doi: 10.1110/ps.5701. [View Article] [PubMed] [Google Scholar]
- Földes-Papp Z, Rigler R (2001) Quantitative two-color fluorescence cross-correlation spectroscopy in the analysis of polymerase chain reaction. Biol Chem 382: 473-478. doi: 10.1515/BC.2001.057. [View Article] [PubMed] [Google Scholar]
- Bark N, Földes-Papp Z, Rigler R (1999) The incipient stage in thrombin-induced fibrin polymerization detected by FCS at the single molecule level. Biochem Biophys Res Commun 260: 35-41. doi: 10.1006/bbrc.1999.0850. [View Article] [PubMed] [Google Scholar]
- Rigler R, Földes-Papp Z, Meyer-Almes FJ, Sammet C, Völcker M, et al. (1998) Fluorescence cross-correlation: a new concept for polymerase chain reaction. J Biotechnol 63: 97-109. [PubMed] [Google Scholar]
- Baumann G, Gryczynski I (2010) Földes-Papp Z (2010) Anomalous behavior in length distributions of 3D random Brownian walks and measured photon count rates within observation volumes of single-molecule trajectories in fluorescence fluctuation microscopy. Optics Express 18: 17883-17896. [PubMed] [Google Scholar]
- Zustiak SP, Nossal R, Sackett . DL (2011) Hindered diffusion in polymeric solutions studied by fluorescence correlation spectroscopy. Biophys J 101: 255-264.[Google Scholar]
- Braet C, Stephan H, Dobbie IM, Togashi DM, Ryder AG, et al. (2007) Mobility and distribution of replication protein A in living cells using fluorescence correlation spectroscopy. Exp Mol Pathol 82: 156-162. doi: 10.1016/j.yexmp.2006.12.008. [View Article] [PubMed] [Google Scholar]
- Feder TJ, Brust-Mascher I, Slattery JP, Baird B, Webb WW (1996) Constrained diffusion or immobile fraction of cell surfaces: a new interpretation. Biophys J 70: 2767-2773. doi: 10.1016/S0006-3495(96)79846-6. [View Article] [PubMed] [Google Scholar]
- Schwille P, Haupts U, Maiti S, Webb WW (1999) Molecular dynamics in living cells observed by fluorescence correlation spectroscopy with one- and two-photon excitation. Biophys J 77: 2251-2265. doi: 10.1016/S0006-3495(99)77065-7. [View Article] [PubMed] [Google Scholar]
- Wu J, Berland KM (2008) Propagators and time-dependent diffusion coefficients for anomalous diffusion. Biophys J 95: 2049-2052. doi: 10.1529/biophysj.107.121608. [View Article] [PubMed] [Google Scholar]
- Dix JA, Verkman AS (2008) Crowding effects on diffusion in solutions and cells. Annu Rev Biophys 37: 247-263. doi: 10.1146/annurev.biophys.37.032807.125824. [View Article] [PubMed] [Google Scholar]
- Szymanski J, Weiss M (2009) Elucidating the origin of anomalous diffusion in crowded fluids. Phys Rev Lett 103: 38102. [PubMed] [Google Scholar]
- Földes-Papp Z (2006) What it means to measure a single molecule in a solution by fluorescence fluctuation spectroscopy. Ex Mol Pathol 80: 209-218.[Google Scholar]
- Földes-Papp Z (2007) True’ single-molecule molecule observations by fluorescence correlation spectroscopy and two-color fluorescence cross-correlation spectroscopy. Exp Mol Pathol 82: 147-155. doi: 10.1016/j.yexmp.2006.12.002. [View Article] [PubMed] [Google Scholar]
- Meroz Y, Sokolov IM, Klafter J (2010) Subdiffusion of mixed origins: when ergodicity and nonergodicity coexist. Phys Rev E Stat Nonlin Soft Matter Phys 81: 10101. [PubMed] [Google Scholar]
- Sokolov IM (2012) Models of anomalous diffusion in crowded environments. Soft Matter 8: 9043-9052.[Google Scholar]
- Weigel AV, Simon B, Tamkun MM, Krapf D (2011) Ergodic and nonergodic processes coexist in the plasma membrane as observed by single-molecule tracking. Proc Natl Acad Sci US.A 108: 6438-6443. doi: 10.1073/pnas.1016325108. [View Article] [PubMed] [Google Scholar]
- Heitzer MD, Wolf IM, Sanchez ER, Witchel SF, DeFranco DB (2007) Glucocorticoid receptor physiology. Rev Endocr Metab Disord 8: 321-330. doi: 10.1007/s11154-007-9059-8. [View Article] [PubMed] [Google Scholar]
- Mikuni S, Tamura M, Kinjo M (2007) Analysis of intranuclear binding process of glucocorticoid receptor using fluorescence correlation spectroscopy. FEBS Lett 581: 389-393. doi: 10.1016/j.febslet.2006.12.038. [View Article] [PubMed] [Google Scholar]
- Cheung J, Smith DF (2000) Molecular chaperone interactions with steroid receptors: an update. Mol Endocrinol 14: 939-946. doi: 10.1210/mend.14.7.0489. [View Article] [PubMed] [Google Scholar]
- McNally JG, Muller WG, Walker D, Wolford R, Hager . GL (2000) The glucocorticoid receptor: rapid exchange with regulatory sites in living cells. Science 287: 1262-1265.[Google Scholar]
- De Bosscher K, Haegeman G (2008) Minireview: latest perspectives on antiinflammatory actions of glucocorticoids. Mol Endocrinol 23: 281-291. doi: 10.1210/me.2008-0283. [View Article] [PubMed] [Google Scholar]
- Vandevyver S, Dejager L, Libert C (2012) On the trail of the glucocorticoid receptor: into the nucleus and back. Traffic 13: 364-374. doi: 10.1111/j.1600-0854.2011.01288.x. [View Article] [PubMed] [Google Scholar]
- Silverman MN, Mukhopadhyay P, Belyavskaya E, Tonelli LH, Revenis BD, et al. (2012) Glucocorticoid receptor dimerization is required for proper recovery of LPS-induced inflammation, sickness behavior and metabolism in mice. Mol Psychiatry 18: 1006-1017. doi: 10.1038/mp.2012.131. [View Article] [PubMed] [Google Scholar]
- Magde D, Elson E, Webb WW (1972) Thermodynamic fluctuations in a reacting system: measurement by fluorescence correlation spectroscopy. Phys Rev Lett 29: 705-708.[Google Scholar]
- Eigen M, Rigler R (1994) Sorting single molecules: application to diagnostics and evolutionary biotechnology. Proc Natl Acad Sci U S A 91: 5740-5747. [PubMed] [Google Scholar]
- Barkai E, Metzler R, Klafter J (2000) From continuous time random walks to the fractional Fokker-Planck equation. Phys Rev E 61: 132-138.[Google Scholar]
- Metzler R, Klafter J (2000) The random walker's guide to anomalous diffusion: a fractal dynamics approach. Phys Rep 339: 1-77.[Google Scholar]
- Havlin S, Ben-Avraham D (1987) Diffusion in disordered media. Adv Phys 36: 695-798.[Google Scholar]
- Gebhardt JCM, Suter DM, Roy R, Zhao ZW, Chapman AR, et al. (2013) Single-molecule imaging of transcription factor binding to DNA in live mammalian cells. Nat Methods 10: 421-426. doi: 10.1038/nmeth.2411. [View Article] [PubMed] [Google Scholar]
- Mazza D, Mueller F. Stasevich TJ, McNally JG (2013) Convergence of chromatin binding estimates in live cells. Nat Methods 10: 691-692.[Google Scholar]
- Kleiman A, Hübner S, Rodriguez Parkitna JM, Neumann A, Hofer S, et al. (2011) Glucocorticoid receptor dimerization is required for survival in septic shock via suppression of interleukin-1 in macrophages. FASEB J 26: 722-729. doi: 10.1096/fj.11-192112. [View Article] [PubMed] [Google Scholar]
- Foldes-Papp Z (2013) Measurements of single molecules in solution and live cells over longer observation times than those currently possible: the meaningful time. Curr Pharm Biotechnol 14: 441-444. [PubMed] [Google Scholar]
- Földes-Papp Z (2005) How the molecule number is correctly quantified in two-color fluorescence cross-correlation spectroscopy: corrections for cross-talk and quenching in experiments. Curr Pharm Biotechnol 6 6: 437-444. [PubMed] [Google Scholar]
- Földes-Papp Z (2007) Fluorescence fluctuation spectroscopic approaches to the study of a single molecule diffusing in solution and a live cell without systemic drift or convection: a theoretical study. Curr Pharm Biotechnol 8: 261-273. [PubMed] [Google Scholar]
- Digman MA, Gratton E (2011) Lessons in fluctuation correlation spectroscopy. Annu Rev Phys Chem 62: 645-668. doi: 10.1146/annurev-physchem-032210-103424. [View Article] [PubMed] [Google Scholar]
- Di Rienzo C, Gratton E, Beltram F, Cardarelli F (2013) Fast spatiotemporal correlation spectroscopy to determine protein lateral diffusion laws in live cell membranes. Proc Natl Acad Sci U S A 110: 12307-12312. doi: 10.1073/pnas.1222097110. [View Article] [PubMed] [Google Scholar]
- Digman MA, Gratton E (2009) Imaging barriers to diffusion by pair correlation functions. Biophys J 97: 665-673. doi: 10.1016/j.bpj.2009.04.048. [View Article] [PubMed] [Google Scholar]
- Cardarelli F, Gratton . E (2010) In vivo imaging of single-molecule translocation through nuclear pore complexes by pair correlation functions. PLoS One 5(5): e10475. doi:10.1371/journal.pone.0010475. [View Article][Google Scholar]
- Földes-Papp Z (2002) Theory of measuring the selfsame single fluorescent molecule in solution suited for studying individual molecular interactions by SPSM-FCS. Pteridines 13: 73-82.[Google Scholar]
- Földes-Papp Z, Baumann G, Demel U, Tilz GP (2004) Counting and behavior of an individual fluorescent molecule without hydrodynamic flow, immobilization, or photon count statistics. Curr Pharm Biotechnol 5: 163-172. [PubMed] [Google Scholar]
